Seminar Up: Effects of Wintertime Hydropeaking on Riparian Zones
Posted by Louis Addo | Seminar


Satu Ramula, an Adjunct Professor in Ecology and Evolutionary Biology from University of Turku, Finland will give a seminar entitled ” The role of soil microbes in plant invasions“. Satu’s areas of expertise are Demographic methods, invasive species, plant ecology, population ecology, and structured population models.
Time and Date: Friday 13th October 2023 from 09:00 to 9:50 CET over zoom (https://kau-se.zoom.us/j/63791052457). You are cordially welcome to join this seminar.

Raviv Gal, PhD student at NRRV, will give a seminar about freshwater mussels as ecosystem engineers. Freshwater mussels are a highly endangered group with a fascinating life history and an important role in the ecosystem. Raviv will tell us about what we know about the role of mussels in freshwaters, with a focus on his own research into their interactions with the rest of the benthic macroinvertebrate community and decomposition processes. Date and Time: Tuesday 19 September 2023 at 13:15 CET. You can join this seminar live on Zoom (https://kau-se.zoom.us/j/63110430909) or in Room 5D306 at Karlstad University campus. You are all invited.
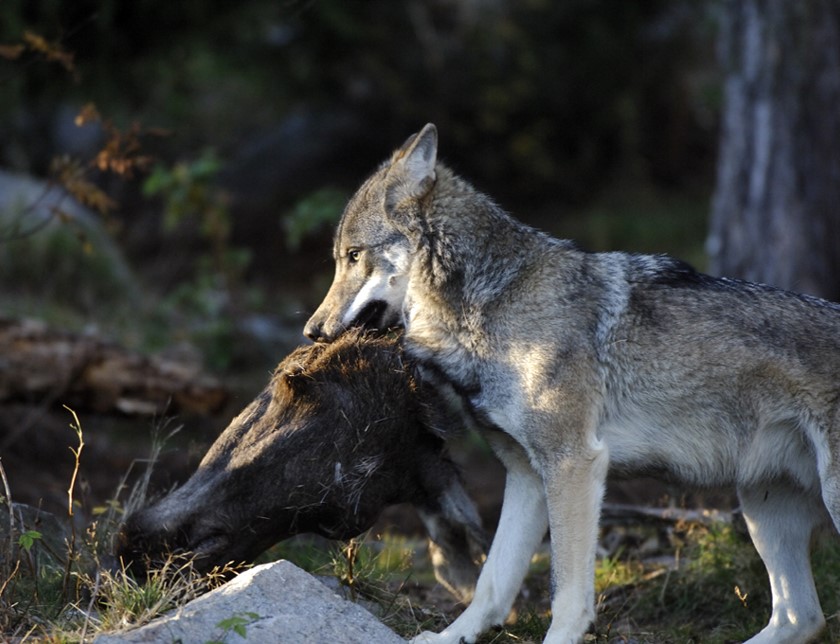

Cecilia Di Bernardi (PhD) from SLU Grimsö will give a seminar about wolf-feeding ecology in Scandinavia and other stories. Date and time: Tuesday 3 October 2023, at 13:15 CET. Join this meeting live over Zoom https://kau-se.zoom.us/j/67521091044 or on Karlsatd University campus at Room 21A349. You are warmly invited.
This post covers chapters 21 & 22 from Futuyma and Kirkpatrick’s book on Evolution (2018). The author of this post is Sam Shry.
Evolution and Homo sapiens; the birds and bees of where we come from
We now turn our attention to the evolution of our own species and how we came to be this unique, evolutionary phenomenon. Of course, we have already learned that we arose from a last universal common ancestor (LUCA), but closer in evolutionary time we arose from the group called great apes, more specifically our closest relatives are the chimpanzee (Pan troglodytes) and the bonobo (Pan paniscus). The lineage hominin (us) diverged from the chimpanzee lineage around 7 Mya, with Homo sapiens becoming the only species from this lineage not to go extinct. With genetic similarities in our protein-coding genes of more than 98% between chimpanzee and humans, it’s no wonder that there are such close similarities between these two species in terms of morphological, social, and cognitive features. We, however, differ in many aspects too, for example, we have evolved the ability to live upright on two feet (bipedal), adapting to a changing climate during this time via movement and migration. We also evolved reduced body hair to probably aid in making sweating and evaporative cooling more efficient. We also have many differences in our hand morphology, teeth, and our larger brain. What is interesting to me is that our potential growth rate is so much higher than for other ape species, as human females can become pregnant even when they have dependent children, in contrast to chimpanzees that cannot.
The divergence of hominins occurred in Africa, as Darwin predicted long before fossil evidence. Hominins split into several species, with all (except us) eventually dying out, giving an interesting insight into the evolution of our species. One of these (Ardipithecus ramidus) had apelike features like a chimpanzee’s brain size and climbing adaptations, but also had small canine teeth and a pelvis adapted for walking upright. The genus Homo dates back about 3 Mya, where the Homo habilis and Homo erectus are thought to be the ancestors of our own species. Homo habilis resembled modern humans closer than earlier species, with a flatter face, shorter tooth row, and greater cranial capacity than before. Homo erectus had a resemblance that was even closer to modern humans and was the first hominin to leave Africa around 2 Mya, spreading into the Middle East and eventually Asia. Our ancestors continued to slowly disperse across the world, with Homo heidelbergensis, a common ancestor to the well-known Neanderthals, dispersing throughout Europe and Asia around 600 Kya (figure 1). I like the analogy in the book that describes the phylogeny of hominins as a densely tangled bush, with humans only being a leaf on this bush.
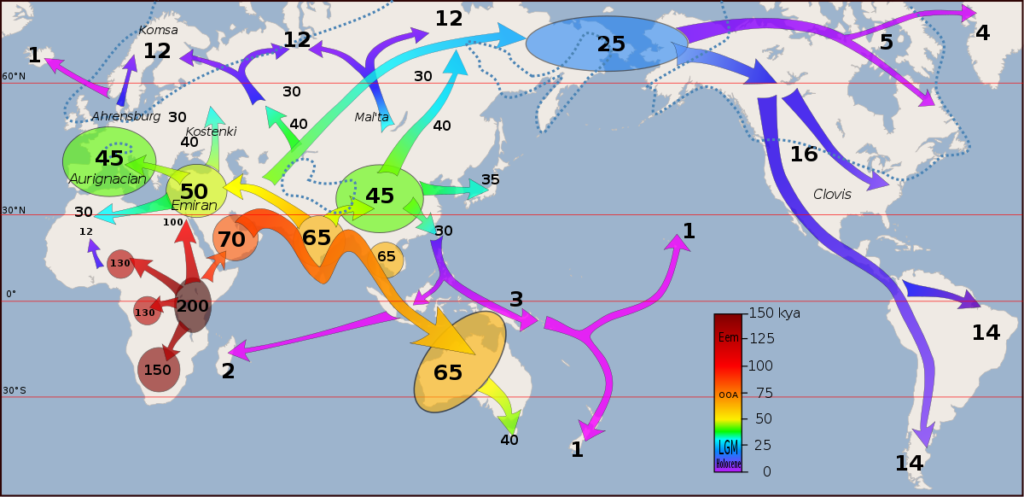
We can map our ancestral dispersal across earth over time using our gene trees of mitochondrial DNA (mtDNA) sequences to complement and provide additional insight to fossil evidence and estimate the age of the most recent common ancestor of mtDNA in living humans. We also find that these different groups of hominins hybridized, for example, Neanderthals and Denisovans, contributing advantageous alleles to the human gene pool. In today’s human populations, we also find some genetic differentiation, but these differences are very low compared to other species, due probably to the very short amount of time we have evolved since colonization. What is surprising is how adapted some human populations are to their environments due to high selective pressure, for example, lighter skin color in Europeans and East Asians adapted to limited sunlight in northern latitudes.
One important characteristic of humans is our brain size. Our cognitive abilities far surpass other species, giving us the ability to reason, think, and understand. One hypothesis for this is ecological, in that we evolved cognition for dealing with complex environments. Another hypothesis is called the social brain hypothesis, where our complex social groups selected for large brains that would ultimately increase survival and the chance for reproduction (increasing fitness). Social cooperation and accumulated knowledge via the use of language are key elements to the evolution of increased cognitive abilities in our species. The evolved modification of our vocal tract enables vocal versatility, enabling a wider range of sounds, but at the cost of possibly choking to death on food, so the advantages of spoken language must be strong in our evolution.
These somewhat strange evolutionary tradeoffs are further seen when looking at the costs of a larger brain, causing higher reproductive costs and requiring more energy to grow all the way up to 18 years, making human life history paradoxical. We do however have longer life spans than other primates, we evolved higher metabolic rates, higher reproductive rates, and a larger energy budget. We did this by switching diets to adding meats and tubers, and learning to hunt and gather. Agriculture was developed and animal domestication began to feed our growing population. Agriculture was double-sided, providing food stability, but changing human lifestyle, culture, and environment around us. Adapting to a new diet selected for specific genes, for example, the lactase persistence enzyme that allows for the digestion of lactose in milk. What is also interesting is how our ancestor dispersal from Africa left traces in our DNA sequence variation, where individuals that disperse furthest from Africa have the highest deleterious mutations, given they were small populations (founder effect), accumulating rare variants, less effective natural selection, and intense drift that fixed these mutations.
Natural selection and evolution are still acting on our species, it’s just that the selection pressures have changed. We find modern hygiene and medicine alleviate the fitness differences, but still, we see selection occurring in some traits, such as cholesterol levels and height, due to our diet, lifestyle, and environmental factors. We also still see the effects of changing to an agricultural diet in our obesity crisis, with calorie-dense foods combined with our sedentary lifestyles. We also have the evolution of culture acting on our society, simultaneously driving cultural elements, innovation, and social interactions. The cumulation of cultural influences, human behavior, and biological tendencies guide, but do not limit our diversity as we continue to move forward in time.
Evolution and Society
The theory of evolution by natural selection has had a profound impact on our understanding of ourselves and the world around us, “unifying the realm of life, meaning and purpose with the realm of space and time, cause and effect, mechanism and physical law.” – Daniel Dennett
Although there is a long line of evidence behind this theory’s confirmation, there are still those that do not accept this theory as reality and cannot grip general reasoning. An extreme, sub-population have “beliefs” in creationism, that God directly created the human species. Certain, literal interpretations of the Bible and other religious documents have conflicting ideas of Earth’s creation and its life, for example in the book of Genesis and the literal interpretation of the creation of Earth and life in six days. Others believe evolution is the mechanism by which God enabled/s creation to proceed and now lets the Earth run on its own without further intervention. People can believe whatever they want, but issues arise when religion and beliefs are confused with science and education. For example, in the US, some schools are required to give equal teaching time to religious doctrine like intelligent design as they do to scientific theory such as evolution. The problem with this is that people do not understand how science works and how it is not comparable to religion. The process of science is to forever test our understanding of the world through the scientific method. Over time we have come up with our best understanding of the world and these denoted theories are “proven” beyond a reasonable doubt via scientific testing and evidence-based science. As this book has explained over the last 21 chapters, the theory of evolution is a highly complex system that we are continuing to understand better every day. Evidence for evolution has been already described throughout the book but we can just recap a few examples. The fossil record and the transitional fossils between species have given us physical evidence of evolution that often matches up, time-wise, with phylogenetic predicted sequences. The phylogenetic relationship between species via DNA analysis has opened a vast new world of understanding every species connection and diversity. Though this evidence does not waver for some true believers, the failed argument of intelligent design gives a baseless ground for these beliefs. The whole argument of a godly design is that these complex species are created perfectly. So, the vast number of failed, inferior designed traits or species in this world would indicate an unkind, incompetent designer. Why would an ultimate designer create sickle-cell disease, saving some and killing everyone else that is homozygous for a gene? Intelligent design is unable to explain the selfish behaviors that natural selection can explain, like cannibalism, siblicide, and infanticide. We have also been able to use the information gleaned from the mechanisms governing the theory of evolution to understand the world around us and how it is changing. For example, understanding domesticated plants and animals, resistance to pesticides and antibiotics, and rapid adaptations to climate changes. In short, the evidence behind evolutionary theory should never be compared to beliefs and even if they were to be, there is no evidence for these beliefs.
Understanding evolution and its mechanisms has shaped our understanding of the world today. It has formed the world we live in, ourselves, and every other species. The concept of natural selection can be applied to all aspects of our lives, such as culture, language, political science, and the overall human experience. The practical application of evolutionary mechanisms has helped us in advancing food production, managing natural resources, improving conservation, and human health. By having this understanding of how organisms evolve, we can make future predictions for our species, other species, and even understand how to develop new technologies and organisms to further benefit us or save our species. This work is vital to combating today’s challenges such as climate change, biodiversity loss, genetic diseases, cancer, infectious diseases, and general public health. One example is the current extinction crises, the global loss of species diversity. Due to rapid climate change, species are unable to genetically adapt to new climate conditions, causing species extinction. Evolution is a unifying theme throughout the biological and social sciences and is the backbone to understanding the world around us. It is, however, only knowledge, with no external meaning, morals, or ethics. It is important knowledge for our society, one species of the millions inhabiting this earth. It is always fun to think about how little evolutionary time our species has inhabited this earth and how/if we will endure or just become another species on the extinction list.
This post covers chapters 18, 19 & 20 from Futuyma and Kirkpatrick’s book on Evolution (2018). The author of this post is Jacqueline Hoppenreijs.
We’ve heard all about the appearing and disappearing of species and species groups on and off the Earth’s surface over the past billions of years…So now it’s time to see how these are linked to the changes that the very surface itself has seen and zoom out a bit, both in space and time!
The geography of evolution
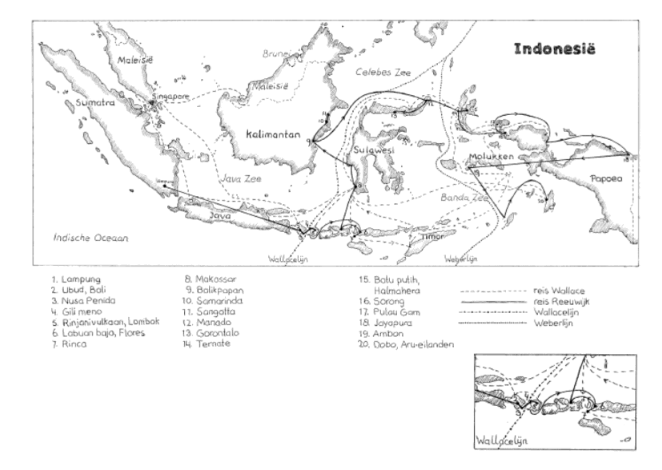
Linking the geographical distribution of species to their ecology is called biogeography, a field that started with the extensive travels of Alexander von Humboldt. He was one of the first to try and figure out why a species occurs in one place but not in another, leading to research on the traits that enable species to live in specific places, measurements of environmental characteristics and, eventually, species’ evolutionary history. Charles Darwin, who was an enormous fan of Von Humboldt (Wulf, 2015), was the first to describe how physical barriers lead to different species occurring in different places, and how similar traits develop if those places are similar. While he was exploring South America, his colleague Alfred Russel Wallace made similar discoveries in south-eastern Asia and Oceania. Following that, Wallace developed a model of biogeographic realms: regions that are inhabited by similar plant and animal taxa. Knowing about the Earth’s geological history, one realises that the current geographical distance between localities is far from the best predictor for ecological similarity of species communities. The realms Wallace himself studied most intensively were the Oriental and the Australian lines, separated by what’s now best known as the Wallace line (Figure 1). The islands of Bali and Lombok, separated by this line, are no more than 50 km apart these days, yet the species groups that inhabit the respective islands are different. The Indomalayan biogeographic realm on the west is maybe best known for its large mammals, such as the Javan rhinoceros and Indian elephant, and its diversity in pitcher plants. On the other side of the divide, the Australian realm is known for marsupials, birds of paradise and eucalypts. There are, however, multiple interpretations of potential lines, of which the Weber line probably is the best-known alternative (Simpson, 1977).
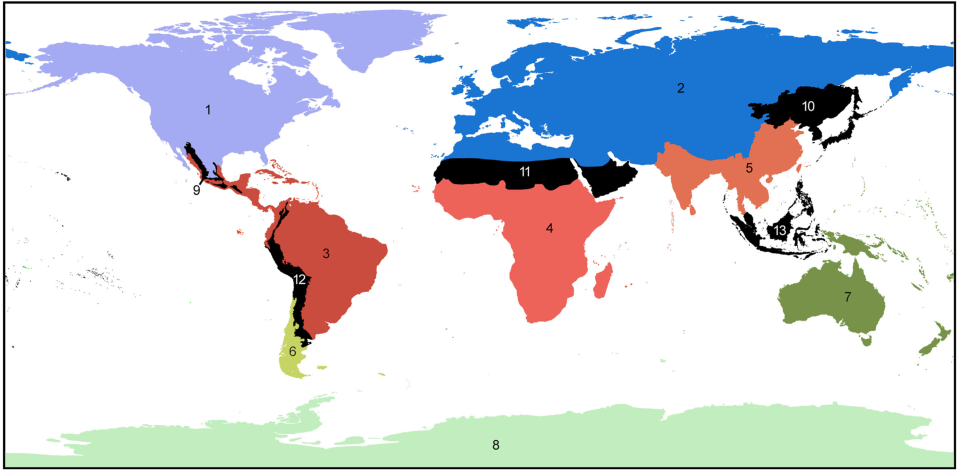
We’ve previously learned about how geographical barriers shape species distributions. While Humboldt, Darwin and Wallace were forced to work with species descriptions and phenotypical measurements, scientists these days have a much wider array of methods at their disposal to map the historical processes of past and current species distributions. The main processes driving these distributions are dispersal, which we have seen can be gradual and look like a stepwise species distribution, and vicariance. Dispersal patterns can, however, also look like giant leaps, for example when a few individuals cover a large distance and start a new population far away from their homes, or when dispersal happened gradually but populations between the two ends of the distribution have been wiped out and gone extinct. This geophysical separation is a form of vicariance, the process that leads to disjunct populations, and often comes in the form of climatological or tectonic disruption. With an increased understanding of plate tectonics and technologies to research fossil remains and build phylogenetic trees, scientists are now able to map the processes driving species distributions better than ever. Our better understanding has led to there being multiple and more nuanced angles to the debate around biorealm delineation, and the general acceptance of there being at least some type of transition zone (Figure 2; Morrone, 2023).
The current distributions of species are a result of these historical processes and current circumstances. The limits of their ranges are decided by several factors, of which dispersal is again an important one: how well is the species able to spread to new areas? That doesn’t only concern the activity itself, but also whether the species finds a mate in this new area (if it depends on sexual reproduction) and how well the species is able to settle in. That means that it needs to be able to find its ecological niche, or in the case of eco-engineers at least to the extent that they can modify it further to fit their requirements. The fundamental ecological niche is the set of abiotic and biotic conditions in which a species finds its most basic needs met, although it might not be the most favourable place. That means that a species A that has its optimum temperature at 17°C but tolerates temperatures between 14 and 24°C, has a niche width of 10°C. Take a species B that thrives at 21°, however, and this species B might outcompete A out of the “upper” part of its fundamental niche. A’s realised niche is thus narrower because of species B, which illustrates the competitive exclusion principle. While this sounds like a relatively modest consequence for species A, it doesn’t sound like a fringe issue at all when you think of the enormous shifts in temperature that the climate crisis causes. Lajeunesse and Fourcade (2023) analysed one of the world’s largest databases, GBIF, on nine different animal taxa and found that almost all of them now consist of species with higher preferred temperatures than 30 years earlier. Since species that are phylogenetically closely related often have similar niches (also called phylogenetic niche conservatism), this means that replacements can be with quite dissimilar species which in its turn can lead to drastic ecological turnovers. Add to the fact that temperature increases are not the only consequence of the climate crisis and the fact that Lajeunesse and Fourcade found that there’s a time lag of these effects too, and we’re looking at a serious problem.
The evolution of biological diversity
While it’s becoming painfully clear that we’re entering the sixth mass extinction (Cowie et al., 2022), it’s not quite as clear how large of an extinction that will be: we simply don’t know how many species inhabit Earth at this time! Current estimations lie between 2 and 8 million eukaryotic species, and we know that those are wildly unevenly distributed in space but also over taxa. It’s not quite clear what drives these differences, but if we’re taking a straightforward approach, we can start by counting species in each of these taxa. If we add information about speciation and extinction rates, we can reason towards a diversification rate, which tells us something about a taxon’s increase or decrease in species richness over time. Two different components of this equation (the time that the taxon has had to undergo diversification and the diversification rate itself), together with the carrying capacity of the system, can help us understand why some species groups are more species-rich than others. A similar argument goes for different areas since areas closer to the equator are usually more species-rich than areas on higher latitudes.
Archaeological records can be used to map past speciation and extinction patterns, and have led to the discovery of, among other things, the Cambrian explosion (which, while sounding violent and destructive, refers to a period of speciation instead of extinction) and previous mass extinctions. The aforementioned sixth mass extinction and its predecessors are not the only time that species or entire taxa became extinct: there is a usual background extinction rate that makes all geological strata potential sources of now-extinct specimens. Both this background extinction rate and the speciation rate have generally declined over the past hundreds of millions of years, and it’s not quite clear why. Especially extinctions should be random, and thus extinction rates quite constant since they depend on random environmental variation and natural selection. Alternative hypotheses are 1) species-rich taxa don’t count as extinct until all of the species within the respective taxa are indeed extinct, and right now we happen to have a lot of species-rich taxa, and 2) the taxa that undergo rapid speciation and extinction are mostly extinct now, and we’re left with the taxa that go through these processes more slowly. The second hypothesis builds on the fact that speciation and extinction rates of taxa are correlated with each other (Figure 3).
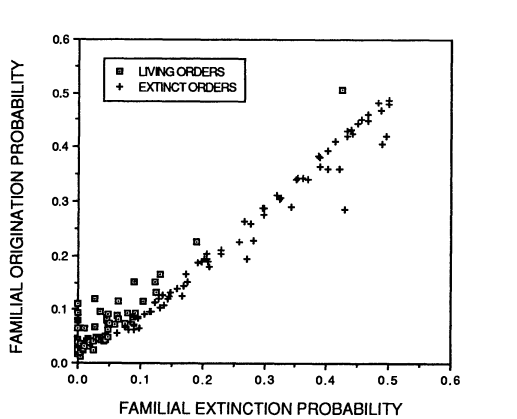
While scientists have found out that extinction rates are relatively constant per taxon, mass extinctions were everything but random: some taxa had better chances to survive than others. Their respective chances depended not on the usual factors, such as consisting of many species, being widely distributed and so on, but on specific characteristic X that allowed them to deal with the cause of mass extinction Y (or not, obviously). One could thus say that evolutionary change works on three levels here: there’s the level of change within species or even populations, the level above on which species originate and go extinct at normal rates a.k.a. species selection, and the final level in which mass extinctions reshape the whole landscape of flora and fauna.
Reshaping consists not only of the wiping out of large parts of species communities, but also offers the opportunity for species to take in the niches that newly opened up and that are relatively competition-free. Mass extinctions are often followed by higher speciation rates, especially if a key adaptation comes to life. Key adaptations are species characteristics that open up a whole new niche, think of the wings of insects that opened up the possibility of flight or lignin in plants that allowed them to build more sturdy structures and colonise land. Such adaptations can lead to increased speciation rates or even periods of radiation, which can result in bouncing back to a certain equilibrium of species richness. Whether such equilibria exist is, however, a topic of discussion, and certainly also depends on the spatial scales one looks at. Local richness can depend on regional richness and is to a certain extent limited by resource availability and competition levels, since higher species diversity tends to lead to lower diversification rates. If not a perfect equilibrium, these dependencies at least suggest a certain bandwidth wherein species numbers fluctuate. If we zoom out, however, we also see how new taxa take in open ecological spaces, suggesting ever-increasing diversity. With that, we can safely assume that there are at least some among us that will have an answer to the sixth mass extinction.
Macroevolution: evolution above the species level
As we have seen for extinctions and speciation, processes don’t always have to be gradual and through small steps. The same seems to be true for evolution: while many changes are small and gradual, there also seem to be sudden developments of characteristics. The field of macroevolution covers these and related questions, such as whether such leaps are as random as small mutations. Evolution above the species level, also called macroevolution, shows surprising parallels with evolution within species (microevolution).
Such parallels include intermediate steps on the level of whole taxa instead of single species; intermediate steps that lead to completely new life forms. Examples are the dinosaur fossils that had feathers long before their carriers were able to fly; these feathers were mainly for insulation. It was only when other characteristics, such as smaller body sizes, changed that we would see animals that started to resemble our modern birds. The same goes for mammals: the skulls that allowed for strong jaws and large brains developed step by step and can, due to the nature of the material, be traced back with the help of fossils.
Such fossils can help us fill the evolutionary gaps between related taxa, for example between whales and other mammals. Now-extinct intermediate species could display different degrees of being amphibious, reduced teeth or moved nostrils, and explain roughly how and when four-legged land mammals went “back” to the sea. Some steps that we know about now seem so large, that some rather call them leaps or saltations. Some researchers have even gone as far as calling the few surviving products of these supposed saltations “hopeful monsters”. Often, however, these monstruous leaps are merely lacking proof of intermediate steps in our fossil record.
All of these steps have something in common. They’ve either been an adaptation to the environment that its carrier could benefit from, or they’ve been a by-product of the adaptation or a random change that didn’t have negative consequences to the extent that they were selected against. The latter may also become beneficial characteristics, with changed circumstances. Adaptations sometimes follow behaviours, such as when species change their feeding behaviour, and can be shaped by modifications without a direct genetic basis during an organism’s development. The complexity of phenotypes that we see today is, to some, enough ground to say that a special someone or something has had a hand in creating all these so perfectly working organisms. Unfortunately for them, our understanding of past steps, both small and (seemingly) large, proves that complexity is no reason to assume creation, rather that complexity might be caused by passing time and adaptive evolution through natural selection. For those that need more convincing, the similar changes that we see evolve independently from each other when taxa adapt to similar living conditions should do the job. Such adaptations are similar in function but clearly have different evolutionary origins, and are called analogous. Their conceptual opposites, homologous adaptations are generally speaking adaptations that have different forms across taxa, but are inherited from a shared ancestor. Biological homology is a bit broader than this, as it includes any characteristic that develops through a similar developmental pathway, i.e. eyes or extremities.
As we now know, the rate of evolution can be wildly different over time, between taxa, and so on. It is measured as an amount of change, for example in a mean trait value, per unit of time. What is tricky about this calculation is that its outcome is usually high when measured on a short time interval, and low when measured over a long time. That’s because of the loss or reversal of newly acquired adaptations, but also because of the previously mentioned niche conservatism.
With so much knowledge about the past under our belt, dare we look into the future? It’s safe to say that the processes that have driven evolutionary change in the past will continue to do so, but their direction and the extent of their respective effects is less clear. It’s possible to discern trends though: changes in a certain direction, that, for example, can be measured through the aforementioned mean trait values. Traits that follow passive trends are equally likely to develop in either direction, whereas an active trend means that development in one direction is more likely than in the other. Trends can become passive or active in a certain direction because of individual selection, speciation or extinction rates and species selection. Two major trends that we can distinguish are those towards higher efficiency and higher complexity. The former can be measured in, for example, the amount of energy it takes to perform certain functions. The extent to which a species needs to be able to fulfil those functions, however, depends on its environment and life cycle, so comparing species and saying that they are better or worse adapted based on a specific trait is quite pointless. The latter, complexity, can be described through hierarchical levels in organisms. Think of our multicellular selves, with our nuclei and mitochondria, our many different types of tissues and the rich bacterial flora that populates our guts, while we are busy being a small part of our human society: certainly a level of complexity that was hard to find when the Earth was purely inhabited by unicellular life. Both levels of complexity and everything in between and beyond are found on our planet today, and there are plenty examples of loss of a kind of complexity in species. Even here goes the adage that it has to be profitable, or at least not too disadvantageous, to maintain complexity when circumstances change.
With these major trends in hand, can we predict what the future will look like? Natural science has shifted from viewing evolution as a kind of progress towards “higher” or “better” organisms (also called orthogenesis) to one where it is everything but a teleological process. That is not to say that there is no predictability at all. After all, we have quite a bit of information about the past and the planet’s status quo. This can help us sketch at least some framework for the nearest future, but it’s safe to say that anything further than that remains a question unanswered for the time being.
References
Cowie, R. H., Bouchet, P., & Fontaine, B. (2022). The Sixth Mass Extinction: fact, fiction or speculation? Biological Reviews, 97(2), 640–663. https://doi.org/10.1111/brv.12816
Futuyma, D., & Kirkpatrick, M. (2018). Evolution (4th ed.). Oxford University Press.
Gilinsky, N. L. (1994). Volatility and the Phanerozoic Decline of Background Extinction Intensity. Paleobiology, 20(4), 445–458.
Lajeunesse, A., & Fourcade, Y. (2022). Temporal analysis of GBIF data reveals the restructuring of communities following climate change. Journal of Animal Ecology, 92(2), 391–402. https://doi.org/10.1111/1365-2656.13854
Morrone, J. J. (2023). Why biogeographical transition zones matter. Journal of Biogeography, April, 1–6. https://doi.org/10.1111/jbi.14632
Reeuwijk, A. (2013). Reizen tussen de lijnen. Dwars door Indonesië met Alfred Russel Wallace. Atlas Contact.
Simpson, G. G. (1977). Too Many Lines; The Limits of the Oriental and Australian Zoogeographic Regions. Proceedings of The American Philosophical Society, 121(2), 107–120.
Wulf, A. (2015). The Invention of Nature: Alexander von Humboldt’s New World. John Murray.

Jacqueline Hoppenreijs, one of the RivEM PhD students, will be giving a seminar entitled Effects of hydrogeomorphology on riparian seedbank and vegetation composition. Join us on 13 June (13:15 CEST) on Zoom via https://kau-se.zoom.us/my/magnuslovenwallerius or in room 5D306 on KAU Campus.
This post covers chapters 16 & 17 from Futuyma & Kirkpatrick’s book on Evolution (2018). The author of this post is Sam Shry.
Phylogeny: how we are all connected
We know that we look different from a crocodile (well, most of us), but how did we diverge from our common ancestors millions of years ago? Can we trace back our common ancestry and how we are related to every other organism on this planet via our genetic relationship and morphological characteristics? That’s what phylogeny tries to understand and with the help of modern genetic analysis, this process has opened our eyes to understanding the phylogenies of every species.
By backtracking each species’ ancestry, we can understand how they are all related and can determine when species split based on morphology, derived features, and genetic architecture. Understanding the exact order of evolution among species can be tricky, as mutations can occur repeatedly, creating sometimes the same evolved characteristics multiple times in multiple phylogenies (homoplasy). One example is the evolution of winged insects. It is a convoluted mess of where and when each species developed or lost wings, making the phylogenetic tree hard to disentangle when basing the entire analysis only on the presence or absence of wings. To avoid these issues, it is important to analyze more than one characteristic when determining relationships and by combining different tools for analysis (morphological differences, genetic differences, etc.). It also depends on the temporal scale; recent evolutionary differences need to be analyzed using faster-evolving parts of the genome, whereas longer time scales require slower-evolving genome sections. What is also important to remember is phylogeny and genealogy of species may not match up completely as groups of genes can be copied and inherited even with speciation, called incomplete lineage sorting. There can also be problems like introgression, where regions of one genome are taken from another species via hybridization, horizontal gene transfer, etc., which can give a misleading picture of the species’ genome. The best way to avoid these problems with differentiating species is to analyze larger regions of the genome to detect more variability between species.
DNA sequencing is done in multiple ways, but one of the most common ways is parsimony, examining how different species are related to one another based on their genetic base changes, simply put, the phylogenetic tree requiring the fewest evolutionary changes. It can, however, do a bad job of parsing the exact phylogeny of where and when individuals split into new species. More robust, statistical methods such as likelihood estimations and Bayesian inference do a better job of differentiating species exactly where and when points of speciation occur. Though, DNA sequencing is difficult when DNA is hard to come by, as with extinct species. This is when old-school methods are still used to construct phylogenies from morphological data instead. In both methods, we are trying to pinpoint the exact time in history when evolutionary changes occurred, so understanding the “when” is an important aspect. We can do this by analyzing the molecular clock within DNA sequences as we have learned about previously, giving us a rate of sequence evolution, which can be used to estimate the time of divergence. Understanding when and where evolutionary changes occurred gives us a road map to exploring reasons for divergence.
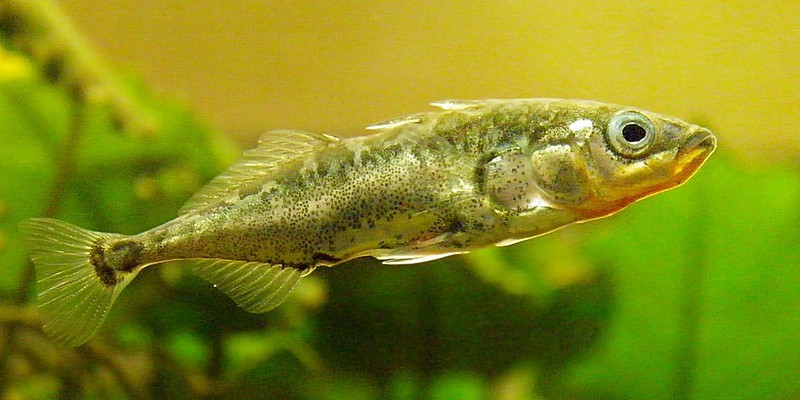
We can use phylogenies to answer questions about mutations, adaptations, and genetic variation in general. One example is the three-spined stickleback, which has invaded freshwater systems of almost all reaches of the Northern Pacific and Atlantic oceans. Their variation in armored plates along their back is quite diverse and has been traced back to a single locus, creating this adaptation throughout freshwater populations. Phylogenies can also be used to trace almost any species’ characteristics, such as language, tails, and even virus prevalence. Phylogenies can be compared to one another, especially when investigating adaptions. Investigating the adaptive coupling of characteristics while controlling for phylogenetic relations is a valuable tool for understanding the evolutionary relationship between species’ characteristics.

Of course, one of the main purposes of developing phylogenies is to classify species into groups and taxa, as has been done from Aristotle, to Linnaeus to Darwin, to modern phylogenetic classifications. Conveying the relationship between species is an important step to understanding biodiversity and leads to further exploration of genetic architecture passed down within and between species.
The history of life… really, that’s the title
A bold title and difficult to summarize into two pages, the underlying connection between today and earth hundreds of millions of years ago are the species that have inhabited this planet over time. We can follow a species through time using their fossils as stepping stones in their evolution, while simultaneously trying to understand how the earth’s environment has changed and impacted species’ biodiversity. As the earth has changed, for example via plate tectonics, the layering or strata of sediment holds records of past organisms and gives us the ability to set a time scale to when organisms were found and when they were lost. These fossils are highly valuable, but very difficult to come by, making our fossil record patchy and incomplete. After the “big bang” 14 billion years ago (Gya), biotic “Life” was formed by abiotic chemical reactions (around 3.5 Gya), creating simple organic molecules. The formation of molecules that could replicate independently (most likely RNA) allowed for evolution by natural selection to begin to shape variation. The complexity of life continued to expand and develop over billions of years, from asexual unicellular to sexual unicellular to multicellular organisms. The Precambrian era (Table 1) brought about evolved photosynthesis and oxygen to the atmosphere, as before this period all life (prokaryotes from the groups Archaea and Bacteria) were anaerobic. The emergence of eukaryotes also occurred (1.8 Gya), opening greater possibilities for diversity.
Table 1. Each step in organic life history and a few important evolutionary points from the millions of years in between.
| Time period | Important stuff that happened |
| Precambrian: 2.5 Gya – 541 Mya | -photosynthesis evolved -oxygen to atmosphere -emergence of eukaryotes |
| Cambrian: 541 Mya – 485 Mya | -explosion of diversity -development of the genetic toolkit for animals -vertebrates arise -ended with large extinction event |
| Paleozoic: 541 Mya – 252 Mya | -species diversity -evidence of modern phyla -movement to land and terrestrial animals -plant diversity and terrestrial modifications -early ancestors of mammals -ended with mass extinction |
| Mesozoic: 252 Mya – 66 Mya | -age of reptiles -birds -break up of Pangea -slow recovery of diversity – adaptive radiation -angiosperms and insect diversity -End with mass extinction via asteroid impact |
| Cenozoic: 66 Mya – today | -modern time -closely related ancestors to modern day organisms -recovery and increased diversity -forming of modern continental geography and climate -adaptive radiation of mammals -human speciation |
The Cambrian era of only about 55 Mya, but created large-scale animal diversity by introducing many new species and classes of animals. This explosion of diversity is attributed to new feeding abilities and novel ways of living. Ecological changes, such as increased atmospheric oxygen and a warmer climate could have also helped in the rise of diversity. More diversity during the Paleozoic period meant an increase in aquatic organism diversity, creating larger and more complex invertebrates and vertebrates. Large squids, sea stars, and molluscs were conceived and some of the first reefs were built by two coral groups. The large extinction event during the late Cambrian gave rise to large-scale diversification with new phyla with new ways of life. Boney fish also arose during this time, creating the ancestors of today’s fish species. Aquatic plant diversity also exploded and advanced to form terrestrial plants with roots that form in the newly acquired terrestrial biomass of organic soil. Insects soon followed, as herbivores feeding on terrestrial plants. The transition from fish to tetrapod to terrestrial vertebrates also occurred during this time, moving from fin to limb. At the end of this era however, a mass extinction event occurred as land masses redistributed themselves, creating chaotic and unfavorable living conditions for many species. The Mesozoic era brought about more stable conditions for living and allowed for yet another round of adaptive radiation and diversification with the rise of new reptile species, both marine and terrestrial. Insects and plants diversified, with angiosperms bringing about insect diversity. There was an explosion in terrestrial insect diversity and the sheer size of some is mindboggling, for example, the Arhropleura, a millipede that reached 2.3m long. Birds also came to be with the diversity in reptiles leading to their wing adaptations for flight. The end of the period is marked by another mass extinction due to the impact of an asteroid off the coast of the Yucatan Peninsula, blocking the sun and creating a long-lasting winter. After that, the Cenozoic era begins around 66 Mya and marks the modern time in evolutionary history. The period is marked by the forming of modern-day continents, climate, and species diversity. The re-diversification formed what are today’s species and the radiation of mammals, eventually leading to the speciation of humans. During this period we also see the first indications of human impact on species diversity, with the extinction of what were called megafauna or large-bodied mammals such as mammoths.
As we can tell from this chapter, life has been ever shaped by the climate and environmental changes on Earth, era after era. Extinctions give rise to adaptive radiations with new species adapting to the present environmental conditions and ever-advancing in complexity. Though today, many of the extinctions occurring are due to human impacts. From our microscopic time span of existence, we are causing rapid global change to both environments and biodiversity. We will just have to see how evolution and Earth sort us out over the next million years…

Larry Greenberg (Professor of Aquatic Ecology, Karlstad University) has been ranked 5313 Best Ecology and Evolution Scientist in the world (https://research.com/scientists-rankings/ecology-and-evolution) and 137 in Sweden (https://research.com/university/ecology-and-evolution/karlstad-university). You can find Larry’s personal Research.com profile here: https://research.com/u/larry-greenberg
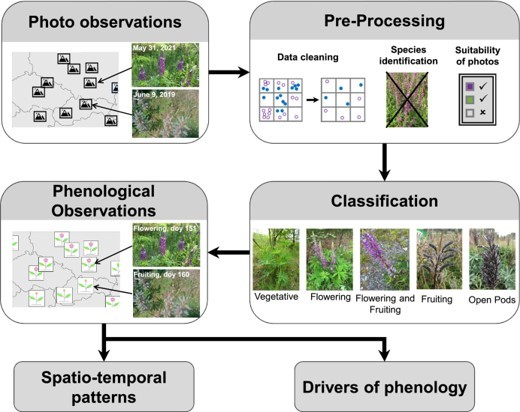
Lutz Eckstein is co-author of a paper presenting a workflow for iPhenology (Fig. 1), i.e., the use of publicly available photo observations to track phenological events at large scales. The paper, which has recently been published open access in the journal Methods in Ecology and Evolution (https://doi.org/10.1111/2041-210X.14114), is led by Yves Klinger (Justus-von-Liebig University Gießen, Germany), who will be visiting Biology at KaU in June 2023. Overall, publicly available citizen science photo observations are suitable to track key phenological events and can thus significantly advance the knowledge on the timing and drivers of plant phenology. In future, integrating the workflow with automated image processing and analysis may enable real-time tracking of plant phenology. To learn more about what sparked the idea for this paper and what may be advantages of working with citizen science photos, see the Methods in Ecology and Evolution blog (https://methodsblog.com/2023/05/22/using-citizen-science-photos-to-perform-phenological-studies/).
… is a research group associated to the subject of Biology and the Department of Environmental and Life Sciences at Karlstad University, Sweden. We conduct both basic and applied research on human impact on river ecosystems, and how this impact can be minimized.
In Swedish the research group is called Naturresurs Rinnande vatten (Acronym NRRV, hence the url nrrv.se).

